[ad_1]
The need for AI and machine learning in decision making across a wide array of industries is no secret. But despite the incredible compute power available, and the constant stream of real-time event data, the subclass of “real-time AI” is somewhat under-addressed, particularly as it pertains to live model retraining and assessment.
Pulsar Functions, a serverless computing framework that runs atop the well-known Apache Pulsar messaging and streaming platform, provides a convenient and powerful solution that addresses limitations in the traditional machine learning workflow. Pulsar Functions takes advantage of the inherent pub/sub nature of the Apache Pulsar platform in order to provide a framework for true real-time AI.
Of course, the scope of Pulsar Functions extends far beyond this domain, but we aim to illustrate the flexibility of Pulsar Functions and how it can serve as a solution for machine learning pipelines that require immediate and real-time predictions and results.
Overview
Our goal is to build a real-time inference engine, powered by Pulsar Functions, that can retrieve low-latency predictions both one at a time and in bulk. To accomplish this, we have two primary development goals:
- Install, configure, and launch Apache Pulsar.
- Define the Python functions that will underpin the inference engine.
The remainder of this article walks through those steps, with a particular focus on the Python development side, as well as the calling interface for both registering and triggering Pulsar functions.
Launching a standalone Apache Pulsar instance
Let’s get started by launching a standalone Apache Pulsar instance. We should note immediately that in a typical deployment of this type of system, Pulsar would be deployed in a cluster, certainly not a standalone instance on a local machine. But a standalone instance will enable us to see the power of real-time AI prior to a more production-quality deployment.
Launch the instance by following the instructions here.
The Pulsar standalone instance will be started with the command:
bin/pulsar standalone
Iris flowers data
With the instance running, we now turn our attention to defining the Pulsar function that will provide our machine learning example. We will use the classic Iris flower dataset. Collected by Edgar Anderson and popularly used by Ronald Fisher, this dataset contains measurements on 50 flowers, spanning three different flower species. The variables included are as follows:
- sepallength: The measured length of the flower’s sepal
- sepalwidth: The measured width of the flower’s sepal
- petallength: The measured length of the flower’s petal
- petalwidth: The measured width of the flower’s petal
- class: The species of flower (Setosa, Versicolor, or Virginica)
A small preview of this data is given below.
|
sepallength |
sepalwidth |
petallength |
petalwidth |
class |
|
5.1 |
3.5 |
1.4 |
0.2 |
Iris-setosa |
|
4.9 |
3.0 |
1.4 |
0.2 |
Iris-setosa |
|
4.7 |
3.2 |
1.3 |
0.2 |
Iris-setosa |
Our goal will be to accurately predict the class given the input features. We will implement the two core components of the model pipeline: training and prediction. The training step will be an offline step (more on this later) while the prediction step will be the core routine of our Pulsar function. Let’s get started!
Building our Pulsar function
Now, we are ready to build and deploy our Pulsar function.
Pulsar functions can be created in a single, standalone Python script, containing the functions which will be deployed. As mentioned, we are going to have the model prediction routine be our primary Pulsar function. A powerful piece of this framework is that the functions themselves are largely standard Python functions, with minimal Pulsar-specific scaffolding. This minimizes the time to deployment for existing code, or the time to creation for more seasoned Python developers.
Let’s start with a basic framework for pulling Iris data, training a model, and writing it to a Pulsar topic.
import os
import pickle
import pandas as pd
from pulsar import Function
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
def train_iris_model():
# If we already have an existing model file, we can load it right away
if os.path.exists("model.pkl"):
print("We are loading a pre-existing model")
return pickle.load(open("model.pkl", 'rb'))
# Read in the iris data, split into a 20% test component
iris = pd.read_csv("https://datahub.io/machine-learning/iris/r/iris.csv")
train, test = train_test_split(iris, test_size=0.2, stratify=iris['class'])
# Get the training data
X_train = train[['sepalwidth', 'sepallength', 'petalwidth', 'petallength']]
y_train = train['class']
# Get the test data
X_test = test[['sepalwidth', 'sepallength', 'petalwidth', 'petallength']]
y_test = test['class']
# Train the model
model = DecisionTreeClassifier(max_depth=3, random_state=1)
model.fit(X_train, y_train)
# Dump the model object to a file
pickle.dump(model, open("model.pkl", 'wb'))
return model
In this code block, we train a decision tree classifier in an attempt to predict the flower species based on the width and length of the sepals and petals. A decision tree classifier can be intuitively represented as a series of decisions based on feature values, culminating in a prediction when a leaf node of the tree is reached. An example tree derived from this model is given below.
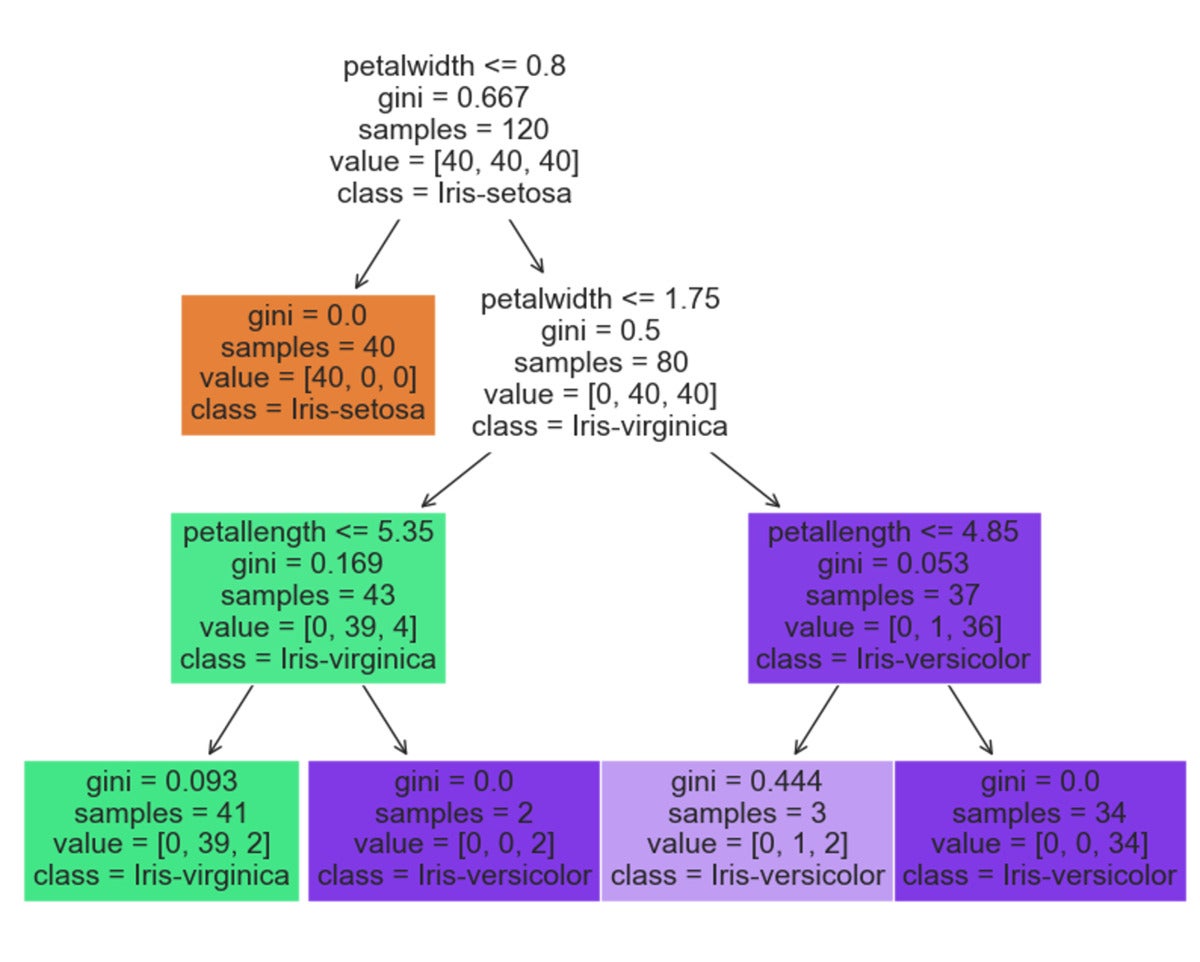 DataStax
DataStaxOne thing to note: We use the pickle module to serialize the model upon training. This will dump the model to a file in the working directory. Subsequent calls to the function will, if the pickled model is available, simply read the model in rather than go through the (sometimes expensive) retraining steps. This will be a key point as we continue fleshing out the example, because it allows for a separate routine that is responsible for continual assessment, augmentation, and re-training of the model as new data is collected.
However, in a production environment, such as when using the Kubernetes runtime for Apache Pulsar, the working directory may be ephemeral, which would instead demand some other solution. A couple possibilities include:
- The model itself can be stored in a Pulsar topic. The serialized pickle format can be written as a base64-encoded string and read on demand rather than reading a file from disk.
- If a physical model file is desired, the pickle file itself can be stored on some remote storage, such as an Amazon S3 bucket. At function run time, the file can be downloaded and read in.
Thus far, we have written the code that will train the decision tree classification model. It’s time to build the routine that will represent our Pulsar function. We will create a subclass IrisPredictionFunction of the Function class in Pulsar, implementing two methods: an __init__() method, which does nothing, and a __process__() method, which, given an input and a user context, returns a prediction from the model.
class IrisPredictionFunction(Function):
# No initialization code needed
def __init__(self):
pass
def process(self, input, context):
# Convert the input ratio to a float, if it isn't already
flower_parameters = [float(x) for x in input.split(",")]
# Get the prediction
model = train_iris_model()
return model.predict([flower_parameters])[0]
This function does not depend on the user context; parameters and configuration options specific to the calling user could be used to tweak the behavior, if desired. But for this function, we simply rely on the input. Because the model was trained on the four flower features sepallength, sepalwidth, petallength, and petalwidth, we must provide each, and in the order that the model was trained. For ease and simplicity, we assume that these are passed as a comma separated string. For example, consider the following string:
1.8,2.1,4.0,1.4
This represents a flower with the following measurements:
- sepallength: 1.8
- sepalwidth: 2.1
- petallength: 4.0
- petalwidth: 1.4
Our Pulsar function will take this string, split on the comma, convert the values to floats, and then pass them to the model prediction routine.
Deploying our Pulsar function
Now for the easy part: deployment! With the Pulsar standalone client running, we need only to create and trigger our function. We first create it, like so:
bin/pulsar-admin functions create \ --tenant public \ --namespace default \ --name iris_prediction_1 \ --py iris_prediction.py \ --timeout-ms 10000 \ --classname iris_prediction.IrisPredictionFunction \ --inputs persistent://public/default/in \ --output persistent://public/default/out
Note a few parameters:
- –name provides us with the name to reference the function to trigger it in the future. This can be whatever is most convenient, but should be unique.
- –py is the name of the Python script which contains the code we have written. Here, I’ve saved our code as a script named iris_prediction.py.
- –classname is the name of the class within the Python script, fully qualified, as shown above.
Finally, we can trigger our function! Let’s pass in the parameters we used as an example above:
bin/pulsar-admin functions trigger \ --tenant public \ --namespace default \ --name iris_prediction_1 \ --trigger-value 1.8,2.1,4.0,1.4
Bonus: Bulk prediction
This function works great for the predicted species of a single Iris flower. But in the real world, we may wish to obtain predictions for a large number of observations at once. Fortunately, this is a trivial modification to the above function, and can be included alongside the single flower prediction function. Let’s add the following class, which we will use to create a new Pulsar function:
class IrisPredictionFunctionBulk(Function):
# No initialization code needed
def __init__(self):
pass
def process(self, input, context):
# Convert the input parameters to floats, if they aren't already
flower_parameters_str = input.split(":")
flower_parameters_split = [x.split(",") for x in flower_parameters_str]
flower_parameters_float = [[float(x) for x in y] for y in flower_parameters_split]
# Get the prediction
model = train_iris_model()
return ", ".join(model.predict(flower_parameters_float))
This bulk version of the function is similar, but differs in three ways:
- It defines a new class name to allow us to distinguish it when registering the function.
- It assumes that we may have more than one set of flower measurements, split by a
:character. - It joins the resulting model predictions into a comma-separated string of predictions, rather than an array.
Much like before, we register the function, making sure to define a new name and refer to the correct class name that we just created:
bin/pulsar-admin functions create \ --tenant public \ --namespace default \ --name iris_prediction_bulk_1 \ --py iris_prediction.py \ --timeout-ms 10000 \ --classname iris_prediction.IrisPredictionFunctionBulk \ --inputs persistent://public/default/in \ --output persistent://public/default/out
And finally, we trigger the function. Let’s pass in three sets of flower measurements at once, like so:
bin/pulsar-admin functions trigger \ --tenant public \ --namespace default \ --name iris_prediction_bulk_1 \ --trigger-value 1.8,2.1,4.0,1.4:0.1,0.1,0.1,0.1:1.8,2.5,0.5,5.0
And there we have it! Real-time predictions of Iris flower species based on the measurements, using Pulsar functions in Python.
While this example merely scratches the surface of what’s possible with Pulsar Functions, I believe it provides a blueprint for implementing a real-time AI pipeline using Apache Pulsar. With Pulsar’s power underlying the core framework, complex pub/sub systems with large amounts of real-time data can be seamlessly processed, with the resulting output from the inference model consumed directly, or even used in a downstream task.
Learn more about real-time AI here.
Eric Hare is a software engineer at DataStax, where he works on open source database, streaming, and AI technologies. Prior to DataStax, Eric held the role of Senior Data Scientist at Daisi Technology. He has a Ph.D. from Iowa State University in statistics and computer science and more than 10 years of experience working with data and analytics.
—
New Tech Forum provides a venue to explore and discuss emerging enterprise technology in unprecedented depth and breadth. The selection is subjective, based on our pick of the technologies we believe to be important and of greatest interest to InfoWorld readers. InfoWorld does not accept marketing collateral for publication and reserves the right to edit all contributed content. Send all inquiries to newtechforum@infoworld.com.
Copyright © 2023 IDG Communications, Inc.
[ad_2]
Source link

